UWI research project to boost Trinidad and Tobago's capacity to monitor virus
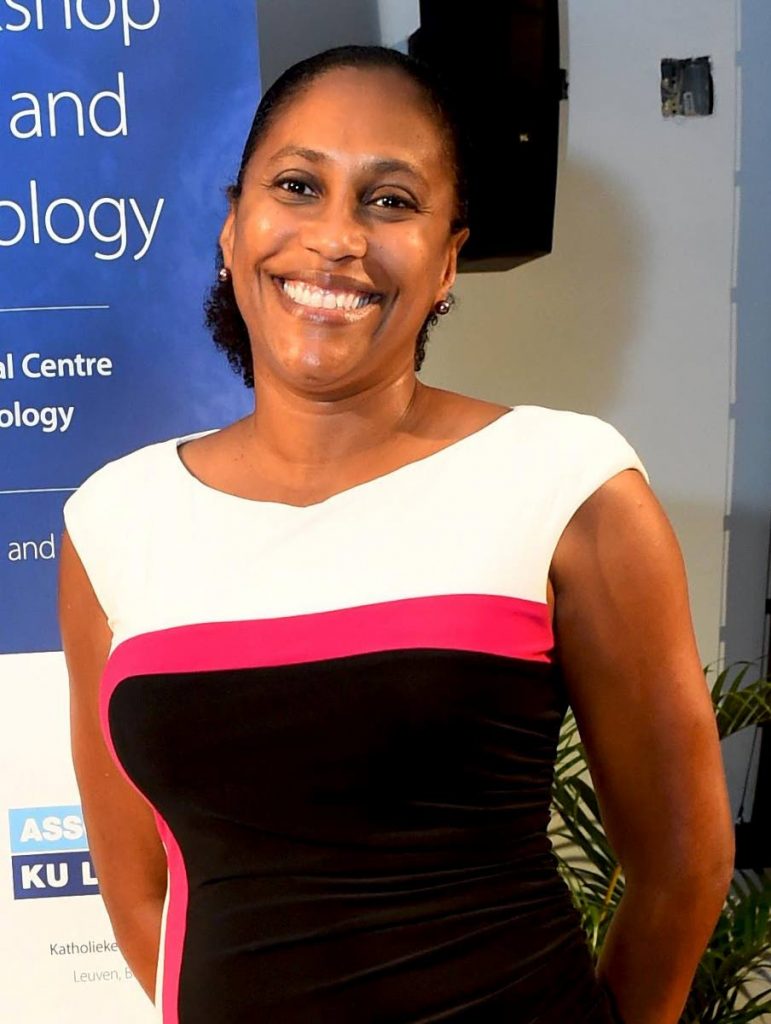

PROFESSOR of molecular genetics and virology at UWI's Faculty of Medical Sciences Christine Carrington says the rapid whole genome-sequencing project she is leading at UWI will be able to address concerns by public health bodies, not just with covid19 but also other infectious diseases in the future.
Genomes are the “hereditary blueprint” of all living organisms, including viruses.
Carrington and a team of researchers from St Augustine campus embarked on the project in December. It involves extracting and sequencing genomes of severe acute respiratory coronavirus 2 (sars-cov-2) from patient samples.
It is titled Covid19: Infectious disease Molecular Epidemiology for Pathogen Control and Tracking (Covid19: Impact).
Carrington said the research will enhance Trinidad and Tobago’s capacity to identify different lineages of the virus and monitor mutations in order to track the virus’s spread, distinguish between local and newly imported cases, and to better understand the virus and the immune system’s response to it.
“The important thing,” Dr Carrington said, “is that whole genome sequencing of the virus can provide epidemiological insights that can inform public health responses. The Nanopore sequencing technology we are using is very rapid. The data is available in real time, so the insights gained are actionable.”
At the Ministry of Health's morning update on January 4, Carrington said her team implemented the project at the request of the Caribbean Public Health Agency (CARPHA). She said the analysis has included that of recent variant sars-cov-2 found in the UK and South Africa, which appears to spread faster than other variants of the virus.
"We analysed virus in diagnostic samples from Jamaica, and showed the presence of the UK variant of concern, which has now been detected in 38 other countries around the world. We have also screened other samples provided by the TT Ministry of Health and by CARPHA for both the UK and South African variants that are of concern, and so far we have not detected them in any of the TT samples. The UWI will continue to support the Ministry of Health and CARPHA with the monitoring."
She said although there are covid19 vaccines on the market, there have been concerns about whether or not they will be effective against the variant strains.
"Will the vaccine protect against it?...While there is growing evidence that the UK and South African (variants) are more transmissible, there is is no evidence that the existing vaccines will not be effective against them, and there is currently no evidence that they are more deadly." But, "From a public health perspective it is important to monitor them and to reinforce the need for non-pharmacological interventions such as wearing masks, hand hygiene and physical distancing."
Carrington said the samples used in the project are from people who were actually tested for covid19, and explained this does not interfere with the diagnostic process.
"When swabs are taken from individuals for diagnostic, the first thing that happens is that they extract genetic material from that sample and that is what they use to do the PCR (polymerase chain reaction) test.
"When they extract that RNA (ribonucleic acid), they don't need all of it for the test, so the excess is stored in a freezer. That's where we get our material."
But because there are insufficient resources to sequence all the samples that are in freezers in TT, she said they just take samples.
"Since we knew these variants of concern (from the UK and South Africa) were in circulation, and we know...that they first appeared around September last year, what we did was look at some of the stored samples as well as some of the more recent samples.
"We will be doing work to look at virus samples from TT and the rest of the region."
She said an analysis of the genetic sequence from the samples will determine the particular lineage that the virus belongs to, from which inferences can be made: "How it is spreading and how lineages are connected and things like that."
The data, Carrington said, has many uses. “You can use it to determine whether a group of cases are linked to each other. For example, if you detect a number of cases in a workplace, you can address questions such as, ‘Are these infections related to each other? Has there been spread within the workplace? Or did each of the affected individuals acquire the infection independently, outside the workplace?’”
This project is funded by a grant from the UWI TT Research, Development and Impact (RDI) Fund, supported by the Ministry of Health, CARPHA, and the universities of Oxford and London.

Carrington said the data will also be very useful when TT has access to a covid19 vaccine.
“We will need to assess the impact of the vaccine. These molecular approaches can be used to monitor levels of viral diversity in the country and to estimate changes in viral population size.”
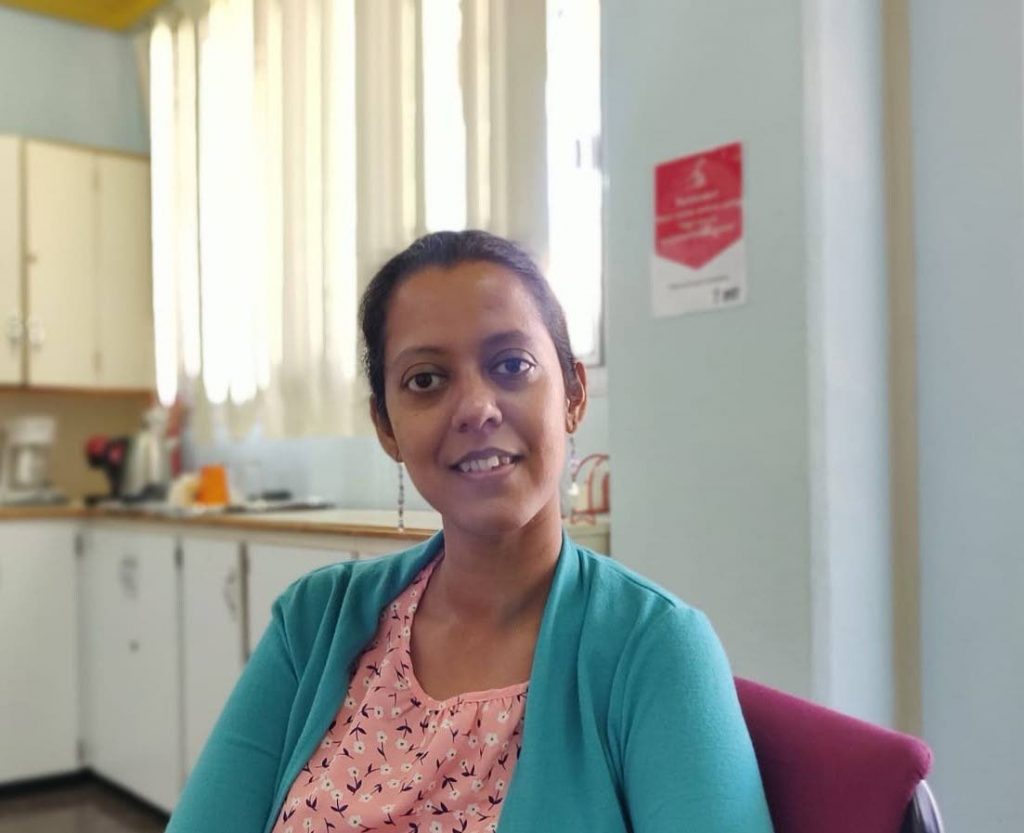
Although the research is still in its early phase, Dr Nikita Sahadeo, post-doctoral researcher on the project, has generated the first five sars-cov-2 genomes from TT. The project will expand, Carrington says, “to get baseline data on the coronavirus lineages that have circulated in TT, the level of diversity we have, and how it has changed over time.
“Once we have that data we will be in a position to better answer questions that public health bodies might have."

Comments
"UWI research project to boost Trinidad and Tobago’s capacity to monitor virus"